|
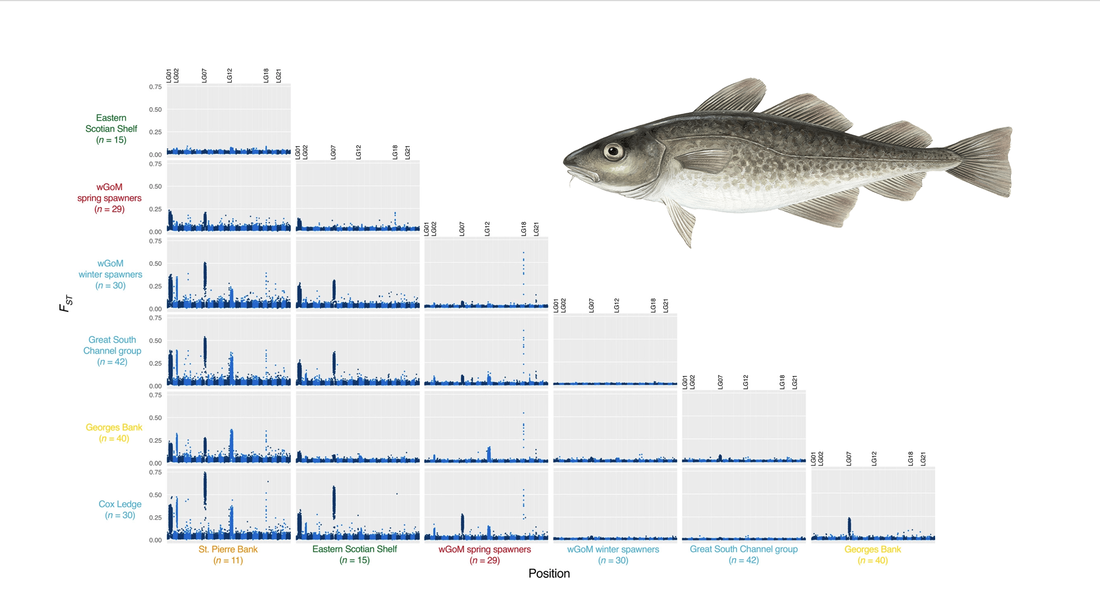
The first results from our collaboration with Gemma Glucas and Adrienne Kovach at the University of New Hampshire has just come out in Evolutionary Applications. See the paper here. We used our very cost-effective low-coverage whole genome sequencing approach to scan for patterns of variation among Atlantic cod sampled from major spawning grounds in the Gulf of Maine and adjacent areas. We saw highly elevated levels of differentiation at four well-characterized large inversions in the cod genome, but our whole-genome dataset also identified multiple narrow peaks of differentiation, the most striking of which differentiate groups of cod that spawn in the same area, but at different times of year (winter vs. spring). Earlier studies based on SNP chips or other more sparse genome sampling would likely have missed these signatures, highlighting a key advantage to sacrificing genotype certainty for the higher resolution afforded by scanning the entire genome through low-coverage sequencing. .The findings of this study clearly show that the current management structure for Atlantic cod does not account for the diversity and population structure in this species, providing important input for Nina's work as part of the NOAA Atlantic Cod Stock Structure Working Group tasked with providing updated scientific advice to managers.
0 Comments
Leave a Reply. |
Therkildsen LabNews updates Archives
April 2023
Categories |

 RSS Feed
RSS Feed